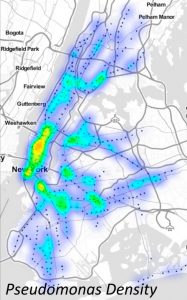
A newly released study has merged big data analytics with the microbiological diversity of New York City’s subways, roughly mapping the DNA of the entire system. The collaborative effort was led by Christopher Mason, Ph.D., of Weill Cornell Medical College. Among the study’s many co-authors was Sergios-Orestis Kolokotronis, Ph.D., assistant professor in the Department of Biological Sciences.
Kolokotronis discussed the research with Inside Fordham last year and described the study as a good way “to understand the bacteria New Yorkers come in contact with on a daily basis,” noting there was surprisingly little data on the subject at the time.
The new study,”Geospatial Resolution of Human and Bacterial Diversity with City-Scale Metagenomics,” has offered a super computer analysis of more than 10 billion fragments of biochemical code.
Among the more salient findings were the presence of bacteria associated with foodstuffs that ranged from mozzarella to chickpeas. At the South Ferry station, ocean submergence from Hurricane Sandy still distinguishes that station from the rest of the system, through the presence of bacteria usually associated with very cold Antarctic environments. To the squeamish, a certain yuck factor predominates with discoveries of fecal matter and rat dander aplenty. But focusing on the repellent aspects misses the point, said Kolokotronis, as “it is to our advantage to have a rich diversity of microbes on and in our bodies.”
“New York City is good system for microbial ecology research because we can study what it means to be in a diverse system,” he said.
Both The Wall Street Journal and The New York Times quoted New York City Department of Health officials as questioning the veracity of the data. Kolokotronis said that he and his colleagues were particularly happy that the paper was published in a new Open Access journal that allows anyone—not just academics—to retrieve the data review it for themselves. He noted that the site, Cell Systems, is “the sister journal to the most established molecular biology journal, Cell.”